User Tools
Sidebar
$\nu$Rnet
Members:
Tony Ahn: Analysis person
Tsung-han Yeh: Parameter person
Bastian Schuetrumpf: Code person
Motivation
It is possible that electron fraction Ye will affect the neutron density resulting in the abundance pattern of the r-process.
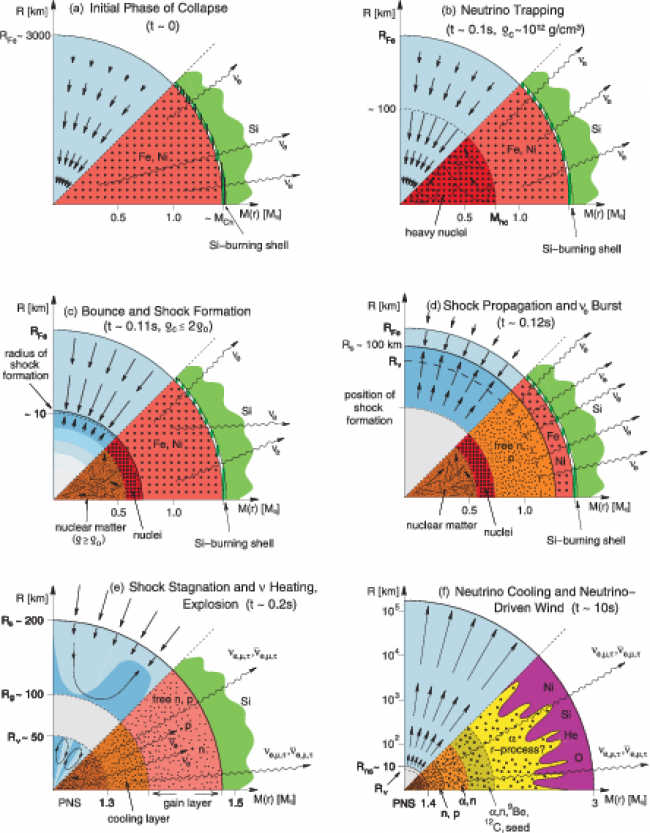
Fig: Schematic representation of the evolutionary stages from stellar core collapse. (Ref. H.Th. Janka, arXiv:astro-ph/0612072)
Method and aims:
- To simulate r-process using x-net
- To change Ye considering effects of the neutrino driven wind
- To find change of the r-process path and change of the abundance pattern
Logbook
Input Parameters
Density temperature trajectory
At first we had to set a reasonable temperature and density trajectory for a neutrino driven wind in which the r-process takes place. We take a realistic trajectory from Raphael Hix the temperature and density profile looks like this:
$Y_e$
To set the initial electron fraction $Y_e$ which is equivalent to the proto fraction due to charge neutrality, we ran the nse version of the xnet code. This calculates the equilibrium initial abundances for a given electron fraction, density and temperature. Therefore the lines in the code tagged by !NSE have to be uncommented and the lines tagged by !NOTNSE have to be commented. This can be done quickly with the following commands in Unix:
sed -e 's/^\!\(.*\!NSE\)/ \1/' conditions.f90 >conditions.f90- sed -e 's/^\!\(.*\!NSE\)/ \1/' net_mpi.f90 >net_mpi.f90- sed -e 's/^\!\(.*\!NSE\)/ \1/' net.f90 >net.f90- sed -e 's/^\!\(.*\!NSE\)/ \1/' data_distribute_mpi.f90 >data_distribute_mpi.f90- mv conditions.f90- conditions.f90 mv net_mpi.f90- net_mpi.f90 mv net.f90- net.f90 mv data_distribute_mpi.f90- data_distribute_mpi.f90 sed -e 's/^\!\(.*\!NOTNSE\)/!\1/' net.f90 >net.f90- mv net.f90- net.f90
Additionally the line
NSE_OBJ = nse.o
in the Makefile has to be uncommented.
It came out that at high Temperatures around 10 GK mainly free neutrons and free protons exist. The other abundances were found to be very low. Thus, for our final calculations we did not use the nse, but changed the proton fraction directly by varying the proton to neutron rate in the initial abundances and neglecting the all other abundances.
Reaction Network
Last but not least we need a reaction library. This was taken from the JINA Reaclib database. For the r-process only nuclides have to be considered which are stable or even more neutron rich. The proton rich nuclides can be neglected. In the chart of nuclides the considered rates look like this:
 The black squares mark the stable isotopes.
The black squares mark the stable isotopes.
This network includes 4510 species and 53763 reaction rates. This setup requires a very fast matrix solver.
Matrix Solver
The standart matrix solver included in the xnet programm “LAPACK” is too slow to solve a network like this. Faster solutions are sparse matrix solver. Two interfaces are included in xnet: For MA48 and Pardiso. Instructions to include these matrix solvers in the code can be found in the documentation of the code. We experienced problems for both with Linux: Pardiso should be much faster than MA48 needs a full compiled LAPACK library. This is included in the ifort compiler. Finally we succeeded in compiling Pardiso version with ifort. The program can be found in the document server.
Results
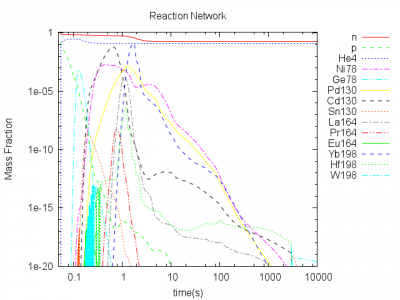
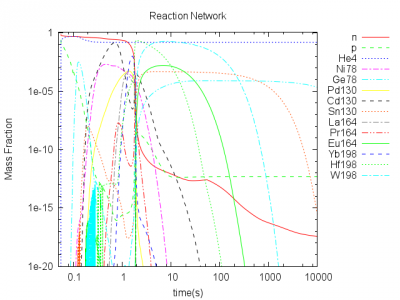
$Y_e=0.15$
$Y_e=0.25$
$Y_e=0.4$
The preliminary results show the abundances of selected nuclides. The abundances are representative for the regions of magic neutron numbers 82 and 126. The abundances in these regions should be high. For $Y_e=0.15$ only small abundances for these nuclides are present, because the seed density consisting of almost the same number of protons and neutrons is too small. For $Y_e=0.25$ we see strong abundances of these nuclides, but for higher $Y_e>0.4$ the neutron density is too small. Thus neutron capure is too slow and the heavy elements cannot be built.